Dallas researchers have developed a new exam based on eight regions of the viral genome that can be easily updated. Implications on the monitoring of variants and on the choice of monoclonal therapies in patients at risk
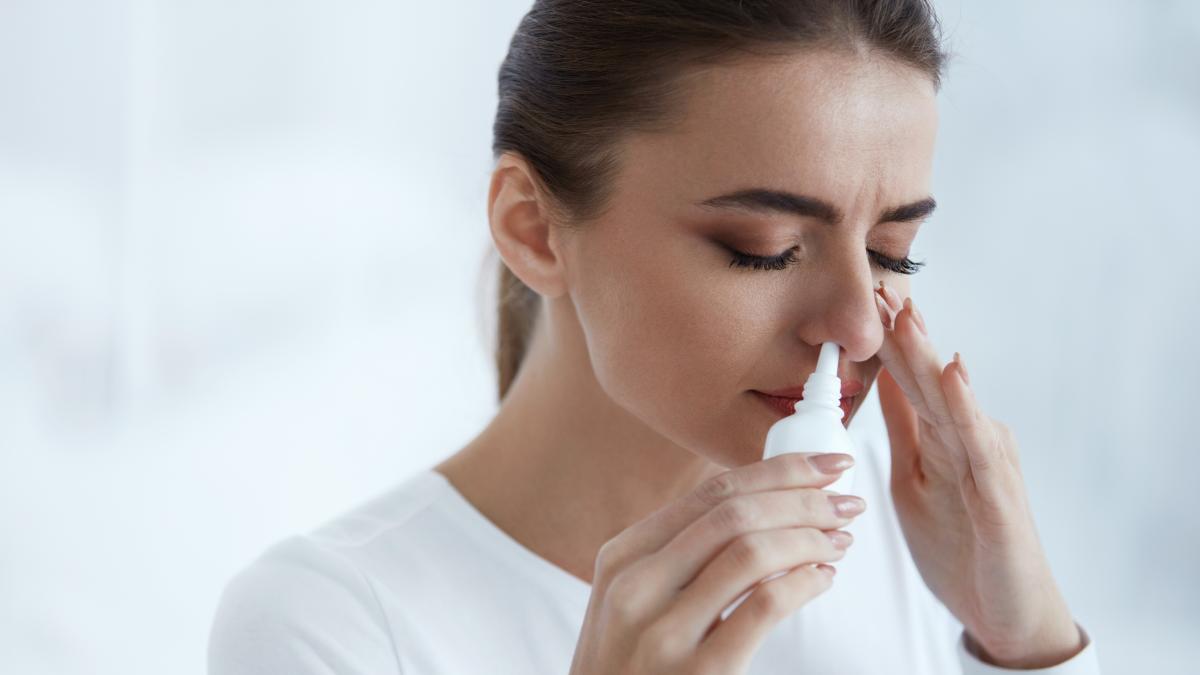
Researchers at UT Southwestern Medical Center in Dallas announced that they have created a test, CoVarScanable not only to detect the positivity to Covid but also to identify the variant.
Using this test, we can very quickly determine which variants are present in the community and if a new variant is emerging explain Jeffrey SoRelle professor of pathology and senior author of the study published in the journal Clinical Chemistry.
According to the researchers, the test could prove useful to monitor the spread of variants and to make clinical decisions quickly, in particular for the choice of monoclonals. The test may be particularly useful on individual patients when we are dealing with variants that respond differently to treatments, adds the researcher.
Pierangelo Clerici, president of the Italian Clinical Microbiologists Association confirms the epidemiological importance of the new test: Quickly knowing the diffusion of the variants is useful if the numbers are important, as now; less if the numbers were to go down. We also don’t know what the cost of the device will be, so at the moment we can’t assess whether the cost-benefits will be real.
There would be no major differences on the choice of therapies for now. Today, the same antivirals are used to combat Covid-19 regardless of the variants. Monoclonals are an exception, and we have seen them work differently depending on the strain. However, if in the future ad hoc therapies are available depending on the variants, the test could be very useful.
How does it work
There are several other tests for Covid-19, but they typically do not provide information to identify the variant. To understand this, scientists have to resort to whole genome sequencinga long and expensive operation that takes a few days.
The CoVarScan focuses on eight regions of the genome of Sars-CoV-2 which differ between viral variants. It detects small mutations – in which the sequence of RNA blocks varies – and measures the length of certain genetic regions that tend to grow or contract with the evolution of the virus. The method relies on polymerase chain reaction (PCR), a common technique in most pathology laboratories, to copy and measure RNA at these eight sites of interest.
I study
To verify the effectiveness of CoVarScan SoRelle performed the tests on over 4,000 positive nasal swab samples to Covid-19 collected between April 2021 and February 2022. The tests were validated with whole genome sequencing, the reference standard and the results were used by doctors to choose treatments in some critically ill Covid-19 patients .
Compared to whole genome sequencing, CoVarScan had a sensitivity of 96% and a specificity of 99%. CoVarScan has identified and differentiated the Delta, Mu, Lambda and Omicron variantsincluding the version BA.2 by Omicron.
The update
But can the new test be easily updated? Responds SoRelle A common criticism of this type of testing that requires constant adjustment for new variants, but the CoVarScan has not needed any adjustments in over a year and continues to perform very well. Meanwhile, BA.2, Omicron 4 and 5 have also appeared and in India a new sub-variant has been identified: BA.2.75 which appears to be even more contagious.
In the future, if we needed to fix it, we could easily add another 20 or 30 hotspots (points in the genome that are sensitive to virus evolution) to the test. Another limitation emerged that the test appears to be highly sensitive. In 95% of the cases tested – explains Pierangelo Clerici – the intercepted viral load was high, with an infectious value, but in a non-negligible percentage the test also identified swabs with a very low viral load, considered non-infectious, as positive. Individuals who cannot infect would also be positive.
July 7, 2022 (change July 7, 2022 | 14:47)
© REPRODUCTION RESERVED
#Covid #developed #rapid #test #identifies #variants #hours